SEAZIT: Systematic Evaluation of the Application of Zebrafish in Toxicology
The small size and rapid development of the zebrafish make it a useful vertebrate model for assessing the potential effects of substances on growth and development using high-throughput screening methods (as reviewed in Planchart et al. 2016). The embryonic zebrafish model has been used for this purpose in pharmaceutical development and in high-throughput screening programs at the NIEHS Division of Translational Toxicology (DTT; formerly the Division of the National Toxicology Program) and EPA. However, broader adoption of the zebrafish model for toxicological screening is hindered by deficits in:
- Consistent experimental protocol elements.
- Clear understanding of mechanisms of chemical absorption, distribution, metabolism, and excretion in zebrafish.
- Consistent classification of developmental toxicity endpoints.
- Consistent informatics approaches used for classification of outcomes.
To promote the broader adoption of zebrafish for toxicological screening, NIEHS DTT established the Systematic Evaluation of the Application of Zebrafish in Toxicology (SEAZIT) program, jointly led by DTT and NICEATM scientists. This webpage summarizes SEAZIT program activities that originated from information gathered from interviews with experts and stakeholders.
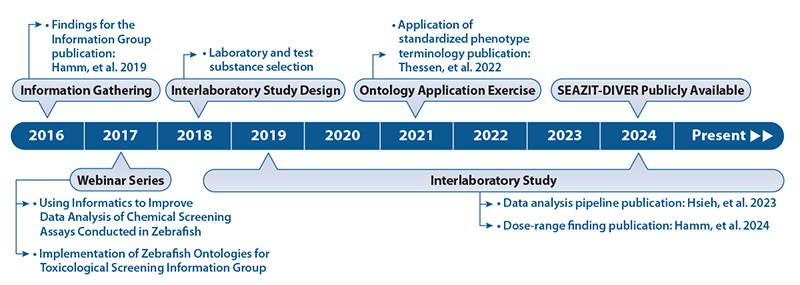
Interviews conducted in 2016 identified areas key to developing a harmonized testing protocol for zebrafish studies, as well as sources of variability among laboratories (Hamm et al. 2019). Following these discussions, the Interlaboratory Study was designed. Deliberations in 2017, summarized in a brief report, focused on the use of ontologies (standardized nomenclature systems) to improve zebrafish screening data analyses. Participants agreed that a comprehensive group analysis of a compiled data set would help demonstrate the advantages of applying a standardized phenotype terminology. As a result, several Collaborative Ontology Efforts were undertaken.
Interlaboratory Study
The interlaboratory study conceived during the Information Gathering phase was completed in 2022.
The study was designed to determine the effect of chorion removal and exposure media renewal on endpoints related to the assessment of developmental toxicity with the embryonic zebrafish model. The initial phase, a dose-range finding (DRF) study, was conducted using the laboratories in-house protocols (Hamm et al. 2024) and helped establish concentration ranges for the second phase of the study as well as assess the laboratories’ capacity to conduct the screening. The next phase, defined as the definitive phase, was conducted using the same chemicals in the DRF but additional requirements for laboratories included the evaluation of test chemicals under four exposure conditions:
- Static exposure of chorionated embryos.
- Static-renewal exposure of chorionated embryos.
- Static exposure of dechorionated embryos.
- Static-renewal exposure of dechorionated embryos.
This study design allowed for evaluating the influence of the chorion and renewal of exposure media within each of the laboratories as well as make comparisons between laboratories to identify other factors that may produce interlaboratory variability. Furthermore, the chemical set, which was designed to provide overlap with historic DTT studies, included chemicals with a range of physicochemical properties and developmental effects. Many of the chemicals have in vivo reference data available from rodent and other zebrafish studies.
A primary goal of SEAZIT is to develop best practices for data analysis. Hsieh et al. 2023 describes various data analysis and reporting strategies utilizing the robust dataset from the interlaboratory study. The data generated in this study can be interactively explored in a web application called SEAZIT-DIVER (Systematic Evaluation of the Application of Zebrafish in Toxicology - Data Integration and Visualization Enabling Resource). SEAZIT data from the interlaboratory study will also be available in the NIEHS Chemical Effects in Biological Systems (CEBS) data resource.
Collaborative Ontology Efforts
Webinar Series
Key issues being addressed by SEAZIT include the variability among laboratories in the endpoints measured and the nomenclature used for each endpoint. On behalf of SEAZIT, NICEATM presented a webinar series in 2017 that considered how these issues might be addressed by implementing ontologies. The goals of the webinar series were to:
- Summarize some of the barriers to routine use of zebrafish in toxicological evaluations.
- Review the concept of ontologies.
- Consider how ontologies could be applied to harmonize procedures used for zebrafish screening studies.
A second webinar series in fall 2018 examined three case studies that illustrated the utility of zebrafish models for toxicology.
Workshop
Following the interviews conducted in 2016, the “Implementation of Zebrafish Ontologies for Toxicological Screening” Information Session was held on April 4 and 5, 2017, in Research Triangle Park, NC. This meeting brought together researchers using zebrafish and data scientists to discuss ontology usage in zebrafish toxicological screening any barriers that exist as well as ways to facilitate future usage. After an overview of current data practices which included several zebrafish researchers sharing their in-house protocols, participants recommended that analysis of a compiled data set should be conducted to demonstrate the advantages of applying ontologies to harmonize zebrafish data. The workshop is summarized in a brief report.
Ontology Application Exercise
To further define ontologies for zebrafish screening, NICEATM coordinated an online evaluation of heterogeneity in terms used to describe zebrafish phenotypes following chemical exposure. Zebrafish images were posted online and researchers asked to evaluate the phenotypes using their in-lab terminology. Results were compiled and terms mapped to the Zebrafish Phenotype Ontology. A second online evaluation employed controlled vocabulary. This study (Thessen et al. 2022) demonstrated that general phenotype terms showed some improvement in consistency between laboratories while granular phenotypes were more difficult to annotate in the second evaluation. This may have been due to raters being given too many choices for granular phenotypes which in turn increased variation. Raters may not have experience with these granular phenotypes suggesting that training should accompany standardization of terminology.
Future Directions
Ancillary projects stemming from SEAZIT activities will be reported on this website when applicable. Please contact Kristen Ryan with any SEAZIT-related questions.


 References:
References: